Sequencing of Catalytic Serine Protease, Linker, and Activation Peptide Domains-Coding Regions of the F9 Gene in Iraqi Hemophilia B Patients
DOI:
https://doi.org/10.32007/jfacmedbaghdad3218Keywords:
c.572G>A, F9 gene, Hemophilia B, Iraq, Sanger sequencingAbstract
Background: Hemophilia B is an X-linked recessive disorder caused by mutations in the F9 gene, causing bleeding tendency predominantly in males. The mutational spectrum of the F9 gene has not been adequately studied in Iraq.
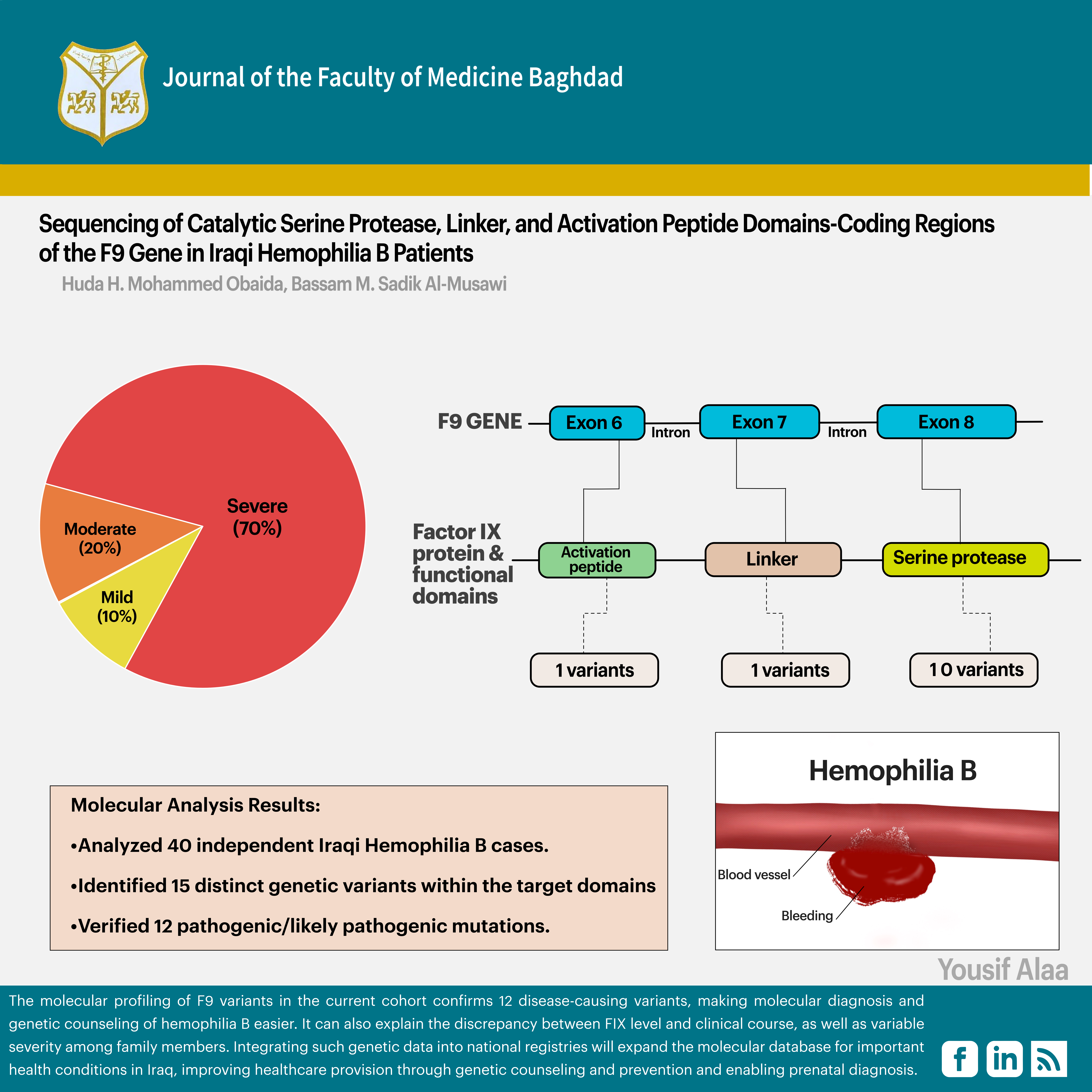
Objectives: To detect the disease-causing variants of exons 6, 7, and 8 and immediate introns of F9 gene using Sanger sequencing among Iraqi hemophilia B patients and to correlate them with phenotypes.
Methods: Forty Iraqi hemophilia B patients were recruited for this cross-sectional study from The Hereditary Bleeding Disorder Ward in the Children Welfare Teaching Hospital, Medical City, Baghdad, between November 2021 and April 2022 using a consecutive sampling technique. Peripheral blood samples were used for sequencing exons 6, 7, and 8, which encode catalytic serine protease (SP), linker, and activation peptide domains and immediate introns of the F9 gene using Sanger sequencing.
Results: Nineteen (47.5%) patients had positive conclusive results. Fifteen unique variants were detected; 12 (80%) of them were disease-causing. Nine variants were located in the SP, one in the linker domain, and two in the splice site of intron 6. The most common pathogenic variant was the c.572G>A (p.Arg191His) on the linker domain as seen in six patients, while c.880C>T (p.Arg294Ter) and c.1358G>T (p.Trp453Leu) were the most common pathogenic variants of the SP domain as seen in two patients each. The vast majority were point mutations that are generally similar to the reported phenotype.
Conclusion: Molecular profiling of F9 gene in the current cohort confirms 12 disease-causing variants, making molecular diagnosis and genetic counseling of hemophilia B possible. It explained the discrepancy between FIX level and clinical course, and variable severity among family members. Integrating genetic data into national registries will expand the molecular database for important health conditions in Iraq, improving healthcare provision through genetic counseling, prevention, and prenatal diagnosis.
Received: Oct. 2026
Revised: Jan. 2026
Accepted: Jan. 2026
Published Online: March 2026
Published: April 2026
References
1.Online Mendelian Inheritance in Man (OMIM). Entry - #306900 - hemophilia B; HEMB. https://www.omim.org/entry/306900/ (accessed 30 August 2025).
2. Kandina DA, Velizhanina ME, Sopova JV, et al. Hemophilia B: Analysis of Mutations in the F9 Gene and Perspectives for the Development of Gene Therapy (A Review). Cell Tiss. Biol. 2025; 19: 291-306. https://doi.org/10.1134/S1990519X25600024 DOI: https://doi.org/10.1134/S1990519X25600024
3. Kaushansky K, Prchal JT, Burns LJ, Lichtman MA, Levi M, Linch DC, editors. Williams Hematology. 10th ed. New York: McGraw-Hill Education; 2021. p.734-738. https://accessmedicine.mhmedical.com/content.aspx?bookid=2962§ionid=252143520/(accessed 2 October 2025).
4. Santoro C, Quintavalle G, Castaman G, et al. Inhibitors in hemophilia B [Internet]. Seminars in Thrombosis and Hemostasis. Thieme Medical Publishers. 2018;44(06):578-589. https://doi.org/10.1055/s-0038-1660817 DOI: https://doi.org/10.1055/s-0038-1660817
5. Castaman G, Motlagh H, Pezeshkpoor B. Hemophilia B: Diagnosis and Management. In: Dorgalaleh A, editor. Congenital Bleeding Disorders. Springer, Cham;2023. https://doi.org/10.1007/978-3-031-43156-2_5 DOI: https://doi.org/10.1007/978-3-031-43156-2_5
6. Miller CH. The Clinical Genetics of Hemophilia B (Factor IX Deficiency). Appl. Clin. Genet. 2021;14:445-454. DOI: https://doi.org/10.2147/TACG.S288256
https://doi.org/10.2147/TACG.S288256 DOI: https://doi.org/10.2147/TACG.S288256
7. National Center for Biotechnology Information. F9 coagulation factor IX - NIH Genetic Testing Registry.https://www.ncbi.nlm.nih.gov/gtr/genes/2158/ (accessed 2 Nov 2025).
8. Shen G, Gao M, Cao Q, et al. The Molecular Basis of FIX Deficiency in Hemophilia B. Int. J. Mol. Sci. 2022 Mar 2;23(5):2762. DOI: https://doi.org/10.3390/ijms23052762
https://doi.org/10.3390/ijms23052762 DOI: https://doi.org/10.3390/ijms23052762
9. Factor IX Variant Database. Factor IX gene (F9). https://www.factorix.org/statistics.html.php/ (accessed 6 September 2025).
10. Hu P, Qiao F, Yuan Y, et al. Noninvasive prenatal diagnosis for X-linked disease by maternal plasma sequencing in a family of Hemophilia B. Taiwan. J. Obstet. Gynecol. 2017 Oct 1;56(5):686-90. DOI: https://doi.org/10.1016/j.tjog.2017.08.020
https://doi.org/10.1016/j.tjog.2017.08.020 DOI: https://doi.org/10.1016/j.tjog.2017.08.020
11. Liu C, Lou X, Lyu J, Wang J, Xu Y. Prenatal Diagnosis and Preimplantation Genetic Diagnosis. In: Pan S, Tang J, editors. Clinical Molecular Diagnostics. Singapore: Springer; 2021. p.769-800. https://doi.org/10.1007/978-981-16-1037-0_43 DOI: https://doi.org/10.1007/978-981-16-1037-0_43
12. Kadhim KA, Al-Lami FH, Baldawi KH. Epidemiological profile of hemophilia in Baghdad-Iraq. Inquiry-j health car. 2019 May; 56:0046958019845280. DOI: https://doi.org/10.1177/0046958019845280
https://doi.org/10.1177/0046958019845280 DOI: https://doi.org/10.1177/0046958019845280
13. Al Rahal NK, Hassan MK, Abid OA, et al. Haemophilia care in Iraq; a multi-centre study. Journal of the Pakistan Medical Association. 2024;74 (Suppl 08): S86-S90. https://doi.org/10.47391/JPMA-BAGH-16-20 DOI: https://doi.org/10.47391/JPMA-BAGH-16-20
14. Centers for Disease Control and Prevention. CDC Hemophilia Mutation Project (Champ & CHBMP).https://www.cdc.gov/hemophilia/mutation-project/index.html/ (accessed 28 Oct 2024).
15. Houge G. Variant classification and reporting. European Society of Human Genetics; 2016. Available from: https://www.eshg.org/fileadmin/www.eshg.org/documents/Variant_classification_system/Variant_class_ESHG.pdf./( accessed 26 April 2025).
16. Lohr SL. Sampling: Design and analysis. 3rd ed. Boca Raton (FL): Chapman and Hall/CRC; 2021 [Internet]. https://doi.org/10.1201/9780429298899 DOI: https://doi.org/10.1201/9780429298899
17. National Center for Biotechnology Information. F9 gene- NLM. https://www.ncbi.nlm.nih.gov/search/all/?term=F9/ (accessed 20 Oct 2025).
18. El-Kamah GY, Mosaad RM, Taher MB, et al. Defining the molecular pathology and consequent phenotypes in Egyptian HB patients. J. Genet. Eng. Biotechnol. 2021 Dec;19(1):1-6. https://doi.org/10.1186/s43141-021-00165-8 DOI: https://doi.org/10.1186/s43141-021-00165-8
19. Parrado Jara YA, Hazbun LKY, Linares A, et al. Molecular characterization of hemophilia B patients in Colombia. Mol. Genet. Genomic Med. 2020;8(5): e1210. https://doi.org/10.1002/mgg3.1210 DOI: https://doi.org/10.1002/mgg3.1210
20. Kulkarni S, Hegde R, Hegde S, et al. Clinical profile of haemophilia B patients from Karnataka. J. Family. Med. Prim. Care. 2022 Jun 1;11(6):2735-2738. https://doi.org/10.4103/jfmpc.jfmpc_1773_21 DOI: https://doi.org/10.4103/jfmpc.jfmpc_1773_21
21. Owaidah T, Bakr S, AbaAlkhail H, et al. Phenotype/genotype screening pattern of hemophilia a and b in saudi arab. Hematol Transfus Cell Ther. 2022 Oct 1;44:S28-9. https://doi.org/10.1016/j.htct.2022.09.1239 DOI: https://doi.org/10.1016/j.htct.2022.09.1239
22. Kizilocak H, Young G. Diagnosis and treatment of hemophilia. Clin. Adv. Hematol. Oncol. 2019 Jun 1;17(6):344-51. Available from: https://www.hematologyandoncology.net/files/2019/06/ho0619KizilocakYoung-1.pdf
23. Header MD, Khaleel MA, Muhabes FJ. Assessment of Mothers' Practices Regarding their Children with Hemophilia in Blood Disease Center at Maternity and Pediatric Babylon Hospital/Al-Hilla City. Kufa. J. nurs. sci. 2016; 6(1). DOI: https://doi.org/10.36321/kjns.vi20161.2613
https://doi.org/10.36321/kjns.vi20161.2613 DOI: https://doi.org/10.36321/kjns.vi20161.2613
24. World Population Review. Inbreeding by Country 2025. https://worldpopulationreview.com/country-rankings/inbreeding-by-country/ (accessed 23 October 2025).
25. Al-Rahal NK. Inherited bleeding disorders in Iraq and consanguineous marriage. Int. J. Hematol. -Oncol. Stem Cell Res. 2018 Oct 1;12(4):273. https://doi.org/10.18502/ijhoscr.v12i4.105 DOI: https://doi.org/10.18502/ijhoscr.v12i4.105
26. Bildirici M, Ersin Ö, Kökdener M. An investigation of hemophilia, consanguineous marriages and economic growth: panel MLP and panel SVR approach. Procedia Econ. Financ. 2016 Jan 1;38:294-307. DOI: https://doi.org/10.1016/S2212-5671(16)30203-9
https://doi.org/10.1016/S2212-5671(16)30203-9 DOI: https://doi.org/10.1016/S2212-5671(16)30203-9
27. Alshaikhli A, Killeen RB, Rokkam VR. Hemophilia B. In: StatPearls [Internet].Treasure Island (FL): StatPearls Publishing; 2025. Available from:https://www.ncbi.nlm.nih.gov/books/NBK560792/.
28. Sahoo T, Naseem S, Ahluwalia J, et al. Inherited bleeding disorders in north Indian children: 14 years' experience from a tertiary care center. Indian. J. Hematol. Blood. Transfus. 2020 Apr;36(2):330-6. https://doi.org/10.1007/s12288-019-01233-3 DOI: https://doi.org/10.1007/s12288-019-01233-3
29. Abood GM, Dehiol RK, Mones HM. Prevalence of Hemophilia and Clinicodemographic Characteristics of Hemophilic Patients Aged ≤ 18 Years in Thi-Qar, Iraq. Global Pediatric Health. 2024;11:1-8. https://doi.org/10.1177/2333794X241280119 DOI: https://doi.org/10.1177/2333794X241280119
30. Yaqoob M, Khan S, Atta S, et al. Current trends of Hepatitis C virus genotypes and associated risk factors in hemophilia patients in Pakistan. Trop. Biomed. 2020 Dec 1;37(4):1000-7. https://doi.org/10.47665/tb.37.4.1000 DOI: https://doi.org/10.47665/tb.37.4.1000
31. VarSome. Varsome: The Human Genomics Community. https://varsome.com/ (accessed 30 March 2025).
32. Li F, He L, Chen G, et al. Variant spectrum of F8 and F9 in hemophilia patients from southern China and 26 novel variants. Frontiers in Genetics. 2023; 14: 1254265. https://doi.org/10.3389/fgene.2023.1254265 DOI: https://doi.org/10.3389/fgene.2023.1254265
33. Laffan MA. Haemophilia and von Willebrand disease. In: Hoffbrand AV, Higgs DR, Keeling DM, Mehta AB, editors. Hoffbrand's Postgraduate Haematology [Internet]. 8th ed. Chichester (UK): John Wiley & Sons Ltd; 2025. p.798-818. https://doi.org/10.1002/9781119706687.ch41/ (accessed 18 Oct 2025). DOI: https://doi.org/10.1002/9781119706687.ch41
34. Raut S, Daas A, Rigsby P, et al. Assay discrepancies using human coagulation factor VIII chromogenic kits: Results from a plasma-derived factor VIII collaborative study (BSP112). Pharmeur Bio Sci Notes. 2023; 2023:1-14. Available from: https://pubmed.ncbi.nlm.nih.gov/37272308/
35. Iannaccaro P, Santoro R, Sottilotta G, et al. Thrombosis in Hemophiliacs with Prothrombotic Molecular Defect. Clinical and Applied Thrombosis / Hemostasis. 2005;11(3):359-360. https://doi.org/10.1177/107602960501100318 DOI: https://doi.org/10.1177/107602960501100318
36. Van den Veyver IB. Skewed X inactivation in X-linked disorders. Semin Reprod Med. 2001 Jun;19(2):183-91. https://doi.org/10.1055/s-2001-15398 DOI: https://doi.org/10.1055/s-2001-15398
37. Miller CH, Bean CJ. Genetic causes of haemophilia in women and girls. Haemophilia. 2021; 27(2): e164-e179. https://doi.org/10.1111/hae.14186 DOI: https://doi.org/10.1111/hae.14186
38. Staber J, Croteau SE, Davis J, et al. The spectrum of bleeding in women and girls with haemophilia B. Haemophilia. 2018 Mar;24(2):180-185 https://doi.org/10.1111/hae.13376 DOI: https://doi.org/10.1111/hae.13376
Downloads
Published
Issue
Section
Categories
License
Copyright (c) 2026 Huda H. Obaida, Bassam M.S. Al-Musawi

This work is licensed under a Creative Commons Attribution 4.0 International License.











 Creative Commons Attribution 4.0 International license..
Creative Commons Attribution 4.0 International license..


